Introduction
Brief Introduction about ncFO.
Ferroptosis is a form of regulated cell death driven by lipid hydroperoxide accumulation. Non-coding RNAs (ncRNAs) are a class of RNA transcripts that cannot encode proteins. Here, we developed a ncRNA–ferroptosis association database (ncFO, http://www.jianglab.cn/ncFO/), which documented the manually curated and predicted ncRNAs involved in the regulation of ferroptosis. Collectively, ncFO recorded 90 experimentally verified entries, and the detailed information of experimentally verified entries included ncRNA name, ncRNA type, disease, species, target, regulation, title, description, and PMID. Then, we downloaded 259 ferroptosis genes from the FerrDb database, including 108 ferroptosis driver genes, 69 ferroptosis suppressor genes, and 111 ferroptosis marker genes. Notably, some genes play diverse roles in the ferroptosis process. Next, we acquired experimentally verified ncRNA–ferroptosis gene associations. Thus, we obtained 3260 predicted entries through regulation. Meanwhile, the ncFO database also introduces online co-expression analysis, survival analysis and differential expression analysis. In summary, ncFO offers a user-friendly web server to search and predict ferroptosis-associated ncRNAs, which might facilitate research on ferroptosis and identify potential targets for cancer therapy.
Why we create ncFO database?
Ferroptosis is a particular type of regulated cell death induced by lipid peroxidation. Triggering the ferroptosis process could effectively kill cancer cells, which might be a powerful strategy for cancer treatment. Non-coding RNAs (ncRNAs), including microRNAs (miRNAs), long non-coding RNAs (lncRNAs), and circular RNAs (circRNAs), have been verified to be involved in the regulation of ferroptosis. An overwhelming number of publications have reported that ncRNAs participate in the regulation of ferroptosis. The scattered information was inconvenient for researchers to characterize the ferroptosis-associated ncRNAs from a comprehensive perspective.
What does ncFO database contain?
The ncFO database documented the manually curated and predicted ncRNA–ferroptosis associations. In addition to the predicted ncRNAs, the ncFO database offers an online analysis of ncRNA in the TCGA dataset.
Main page
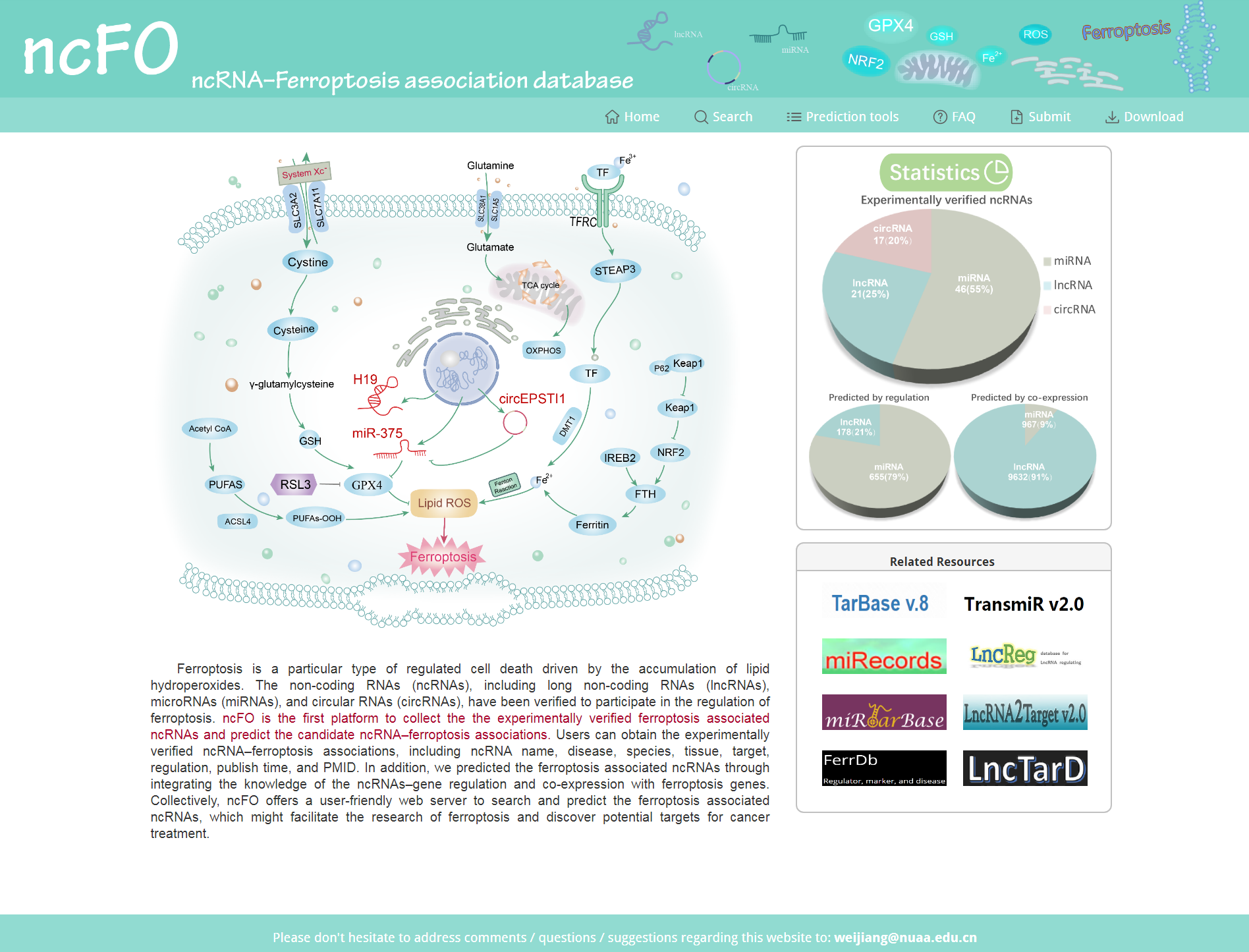
Figure 1. Main page of ncFO.
Search
Search the experimentally validated entries.
Users can search all the experimentally verified entries by ferroptosis genes and three kinds of ncRNAs (miRNA, lncRNA, circRNA). We offer a fuzzy search to recommend all matched miRNA/lncRNA/circRNA lists.
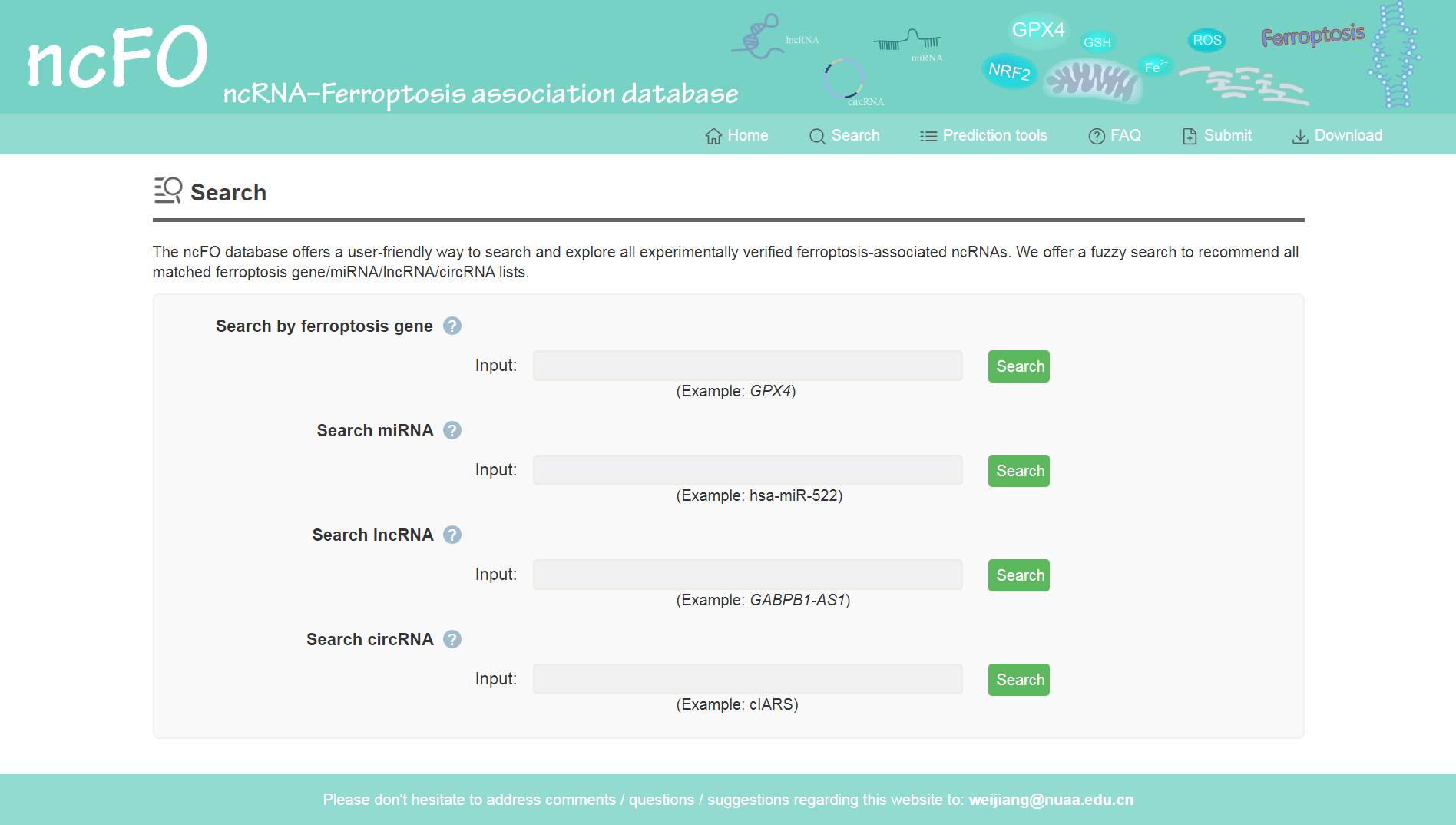
Figure 2. Search the experimentally validated ncRNA–ferroptosis associations.
Search the predicted entries.
Users can predict ferroptosis-associated ncRNAs using two different methods. First, users can search by ferroptosis genes, lncRNAs, or miRNAs to obtain miRNAs/lncRNAs that regulate ferroptosis genes. We also offer a fuzzy search to return all matched ferroptosis gene/miRNA/lncRNA lists. The search results contain basic information, analysis tools, regulatory networks and detailed information of regulations. Second, users could predict ferroptosis-associated ncRNAs using co-expression analysis. By entering the ferroptosis gene, ncRNA type, cutoff, method and specify the cancer type. Then, ncFO returns the correlated ncRNAs based on the input parameters. We implemented R language for online analysis of the ncRNAs in the TCGA dataset.
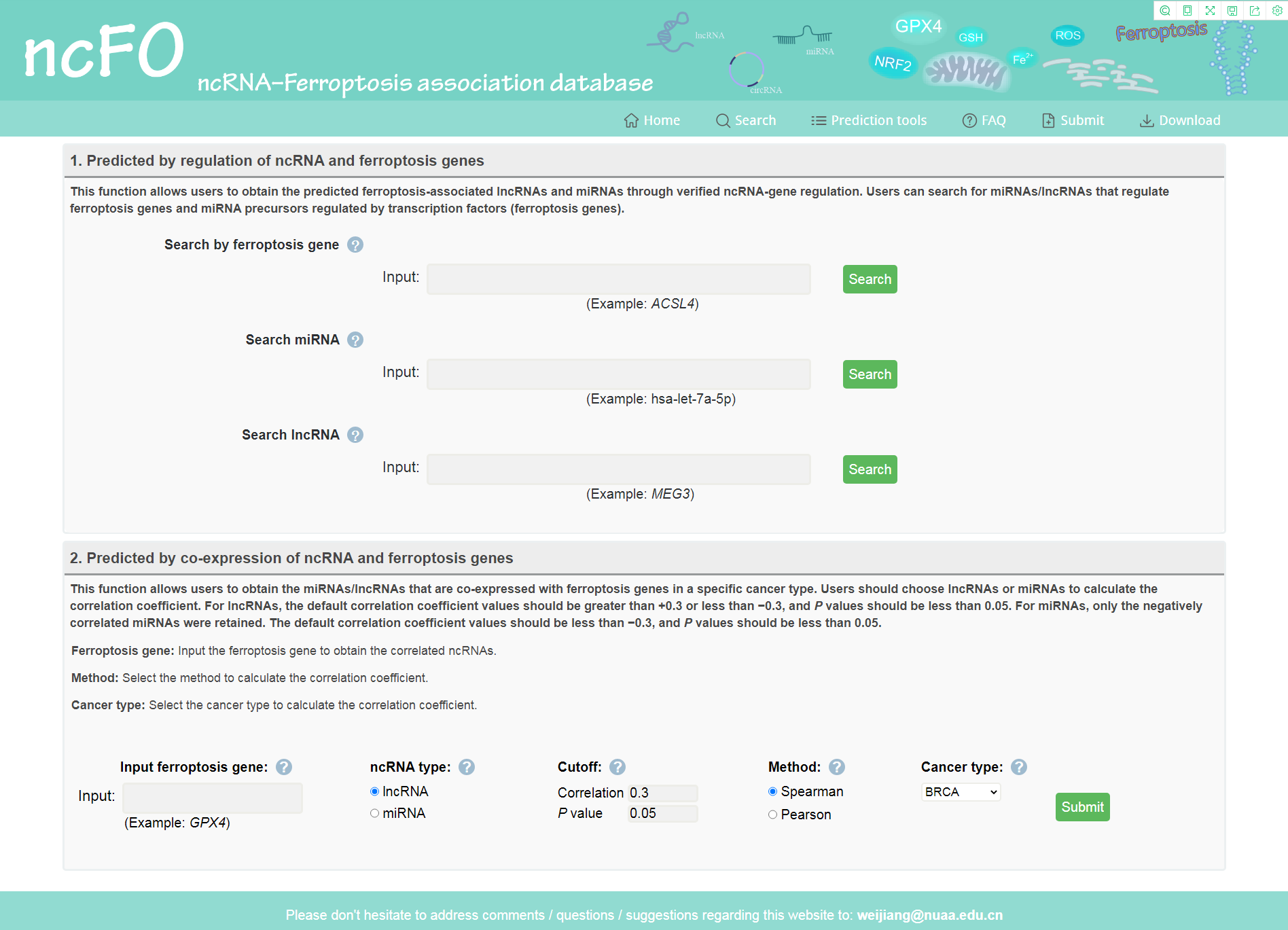
Figure 3. Search the predicted ncRNA–ferroptosis associations.
Search result of "Predicted by regulation".
The search result of "Predicted by regulation of ncRNA and ferroptosis gene", which contains basic information, analysis tools, regulatory networks, and detailed information of regulations. Additionally, users could perform online correlation analysis for the query ncRNA and target genes. Click more to browse the detailed information of the ncRNA-target gene regulation.
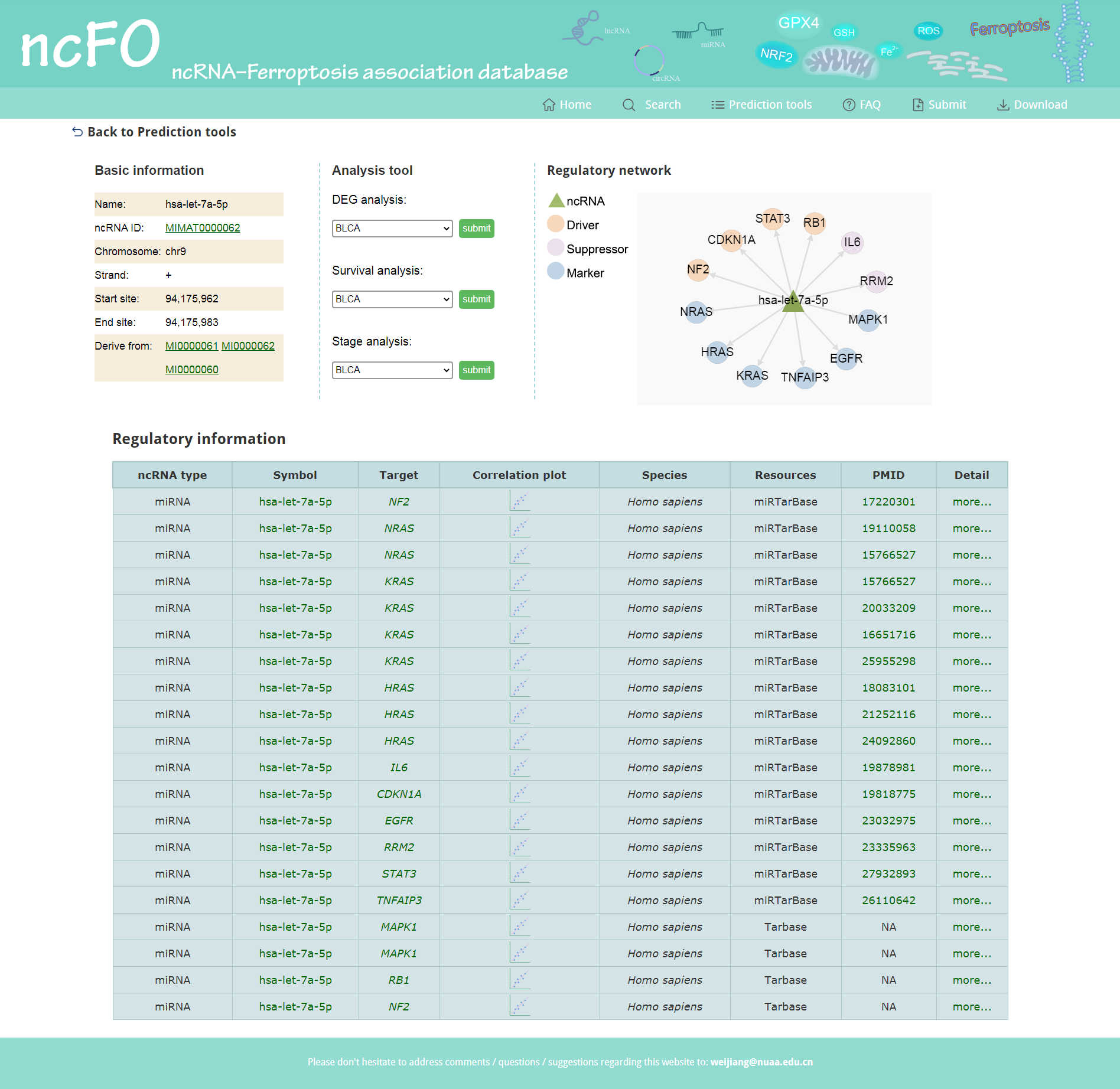
Figure 4. Search result of "Predicted by regulation".
Search result of "Predicted by co-expression".
For the search result "Predicted by co-expression of ncRNA and ferroptosis gene", we calculated the co-expressed ncRNAs of ferroptosis gene. Additionally, we implemented R language for online analysis of the ncRNAs in the TCGA dataset.
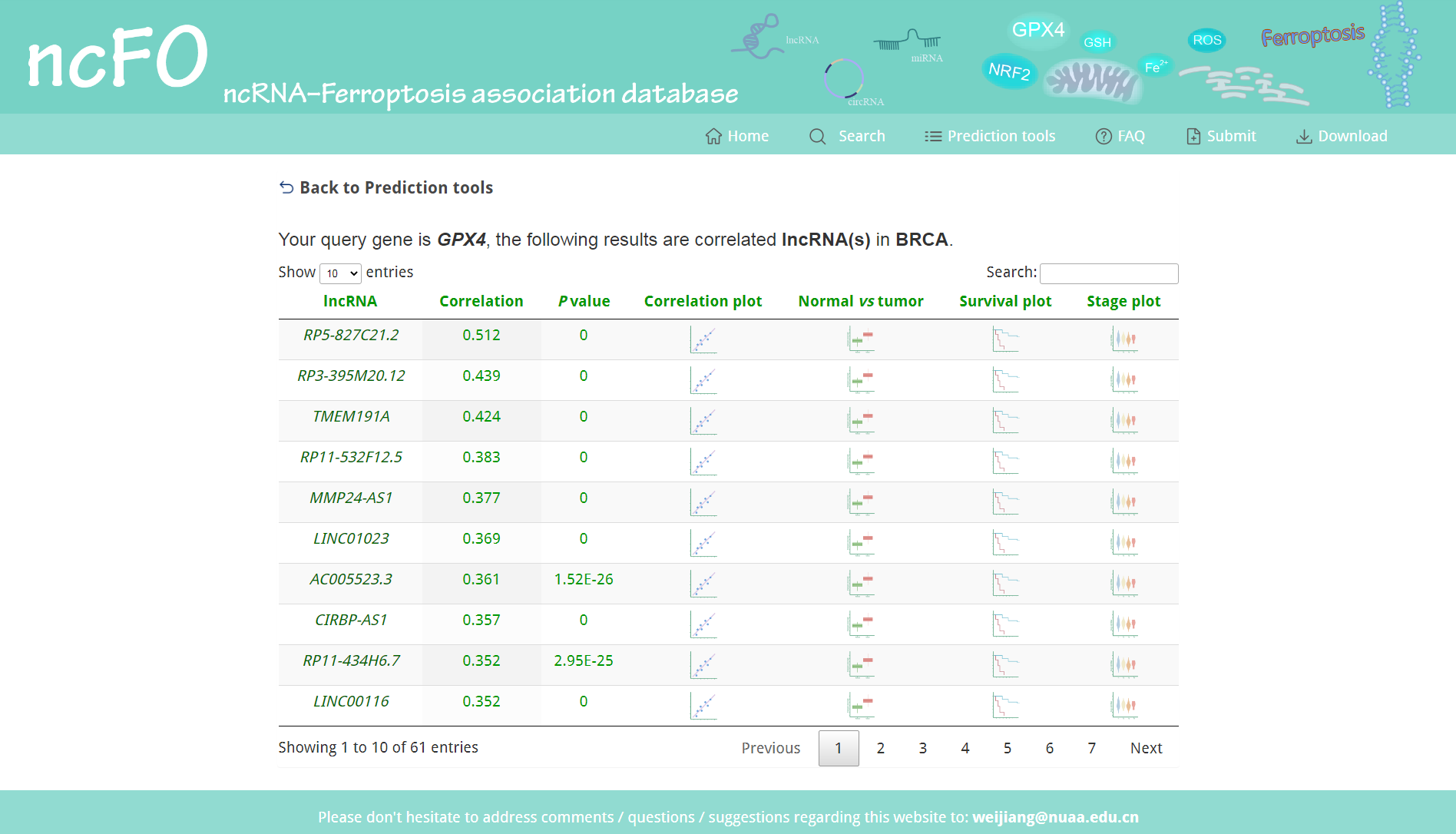
Figure 5. Search result of "Predicted by co-expression".
Submit
Submit new ferroptosis-related ncRNA.
ncFO strongly encourages users to submit novel experimentally supported ncRNA–ferroptosis associations. After inspection of the submitted data, we will update the ncFO database.
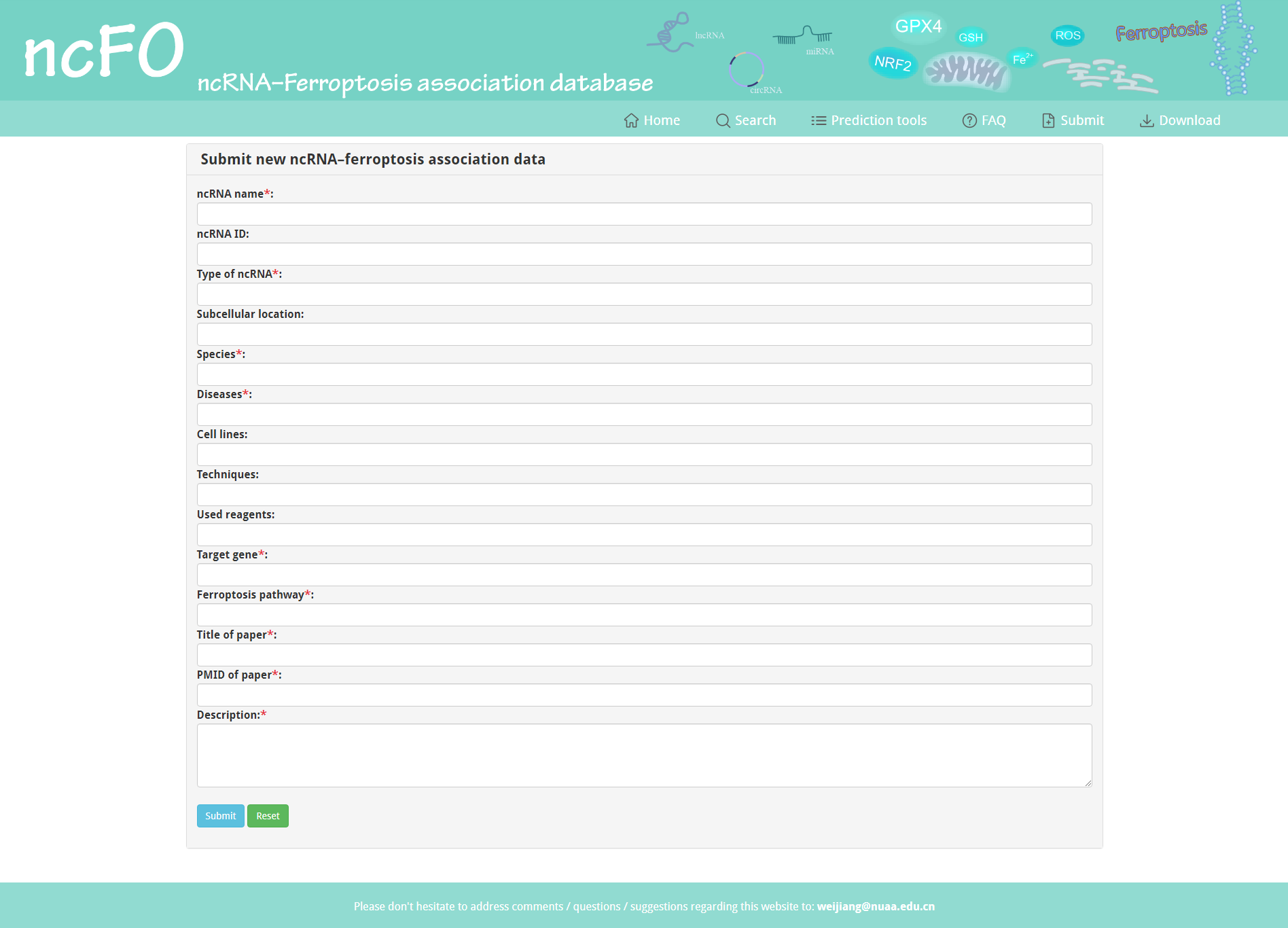
Figure 6. Submit page of ncFO.
Download the ferroptosis-related ncRNAs.
Users could download all of the experimentally verified ncRNA–ferroptosis associations from the ncFO database. In addition, users could download all of the regulation predicted entries, as well as co-expression predicted entries in cancer types.
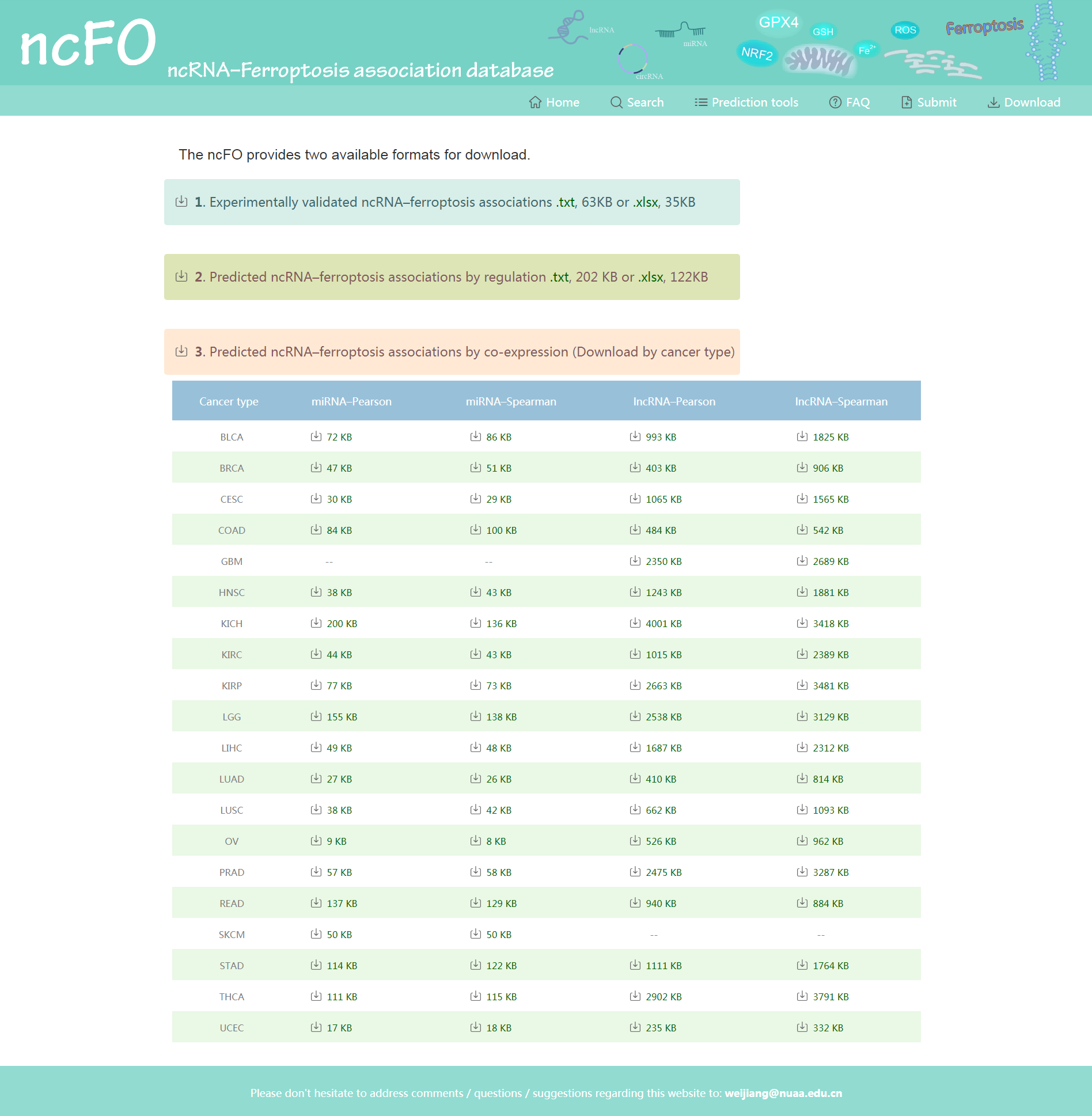

Figure 7. Download page of ncFO.
